Researchers deciphering the mechanism of sliding of DNA clamps, central players in DNA replication and repair
|The genetic information is encoded in long chains of deoxyribonucleic acid (DNA) molecules packaged into chromosomes in the cell nucleus. Every time a cell replicates itself, the DNA needs to be duplicated. Critical players in this process are the so-called DNA clamps, ring-shaped proteins that slide onto the DNA double helix and anchor the polymerases, the enzymes that replicate DNA, onto the genomic template. While two-decade studies revealed the importance of DNA sliding clamps in many cellular processes, key mechanisms of their function at the molecular level remained elusive.
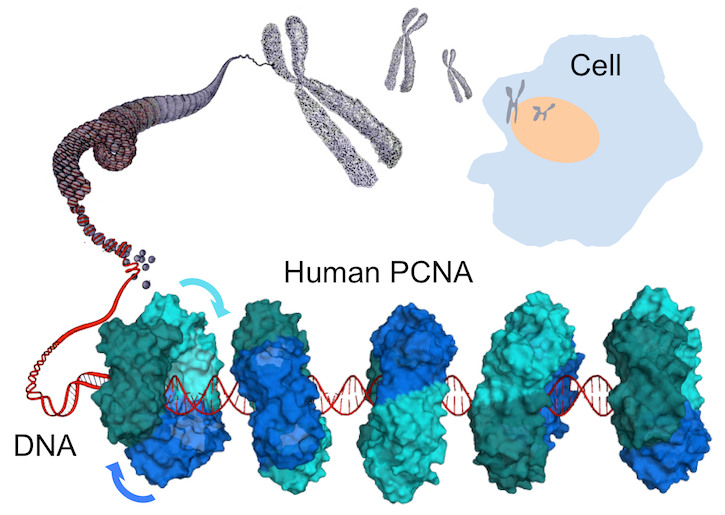
A joint collaboration involving three research teams, and headed by the Structural Biology Laboratory at Elettra-Sincrotrone Trieste (Matteo De March, Silvia Onesti and Alfredo De Biasio), revealed how the human DNA clamp PCNA (Proliferating Cell Nuclear Antigen) moves on DNA. Employing a combination of structural and computational approaches, and also making use of the X-ray beamline XRD1 at Elettra as a CERIC facility, De Biasio and colleagues discovered that PCNA slides on DNA through a spiral motion that keeps the orientation of PCNA competent for binding to the polymerase.
Due to the critical role of DNA sliding clamps in cell proliferation, understanding their mechanics at the molecular level may contribute to defining novel drug targets for anti-cancer therapy. Therefore, the insight provided by this study is also important for potential future medical applications.



